Covariance-Free Sparse Bayesian Learning
Authors
Alexander Lin, Andrew H. Song, Berkin Bilgic, Demba Ba
Abstract
Sparse Bayesian learning (SBL) is a powerful framework for tackling the sparse coding problem while also providing uncertainty quantification. The most popular inference algorithms for SBL exhibit prohibitively large computational costs for high-dimensional problems due to the need to maintain a large covariance matrix. To resolve this issue, we introduce a new method for accelerating SBL inference -- named covariance-free expectation maximization (CoFEM) -- that avoids explicit computation of the covariance matrix. CoFEM solves multiple linear systems to obtain unbiased estimates of the posterior statistics needed by SBL. This is accomplished by exploiting innovations from numerical linear algebra such as preconditioned conjugate gradient and a little-known diagonal estimation rule. For a large class of compressed sensing matrices, we provide theoretical justifications for why our method scales well in high-dimensional settings. Through simulations, we show that CoFEM can be up to thousands of times faster than existing baselines without sacrificing coding accuracy. Through applications to calcium imaging deconvolution and multi-contrast MRI reconstruction, we show that CoFEM enables SBL to tractably tackle high-dimensional sparse coding problems of practical interest.
Concepts
The Big Picture
Imagine tracking a needle in a haystack. Not just locating it, but quantifying exactly how uncertain you are about its position. That’s the challenge behind sparse Bayesian learning (SBL), a technique for reconstructing a complete signal from incomplete, noisy data while accounting for uncertainty.
SBL appears across neuroscience and imaging: reconstructing brain activity in MRI, inferring neural firing from calcium imaging, estimating radio signal directions. Unlike methods like LASSO, SBL doesn’t just return a single answer. It returns a full probability distribution over all possible answers. That honest accounting of uncertainty matters when decisions depend on the result.
The problem? SBL has a computational bottleneck baked into its core. Every update step requires working with a covariance matrix, a grid tracking how every unknown in your signal relates to every other. For D unknowns, this grid has D² entries and requires O(D³) operations to manipulate.
Double the problem size and computation becomes eight times slower. For a brain scan with tens of thousands of voxels, SBL grinds to a halt.
A team from Harvard and MIT built a fix: CoFEM.
Key Insight: CoFEM eliminates the covariance matrix bottleneck by estimating the statistics SBL needs through randomized numerical methods. It matches the accuracy of full-covariance SBL while running up to thousands of times faster.
How It Works
Standard SBL uses an expectation-maximization (EM) algorithm, a two-step cycle that alternates between estimating the signal given current model parameters (the E-step) and updating those parameters (the M-step). The E-step is the bottleneck: it requires forming and inverting a D × D covariance matrix. At D = 10,000, that’s 100 million entries. At D = 100,000, it’s effectively impossible.
CoFEM’s core insight: you don’t need the whole matrix. You only need two quantities from it, the posterior mean (your best single-point estimate of the signal) and the diagonal of the covariance (a compact list of per-unknown uncertainties). Both can be estimated without ever constructing the full matrix.
Here’s how:
-
Solve linear systems instead of inverting matrices. Rather than directly computing Σ = (Φᵀβ Φ + diag{α})⁻¹, CoFEM sets up K linear systems, each with a randomly drawn right-hand side, and solves them using preconditioned conjugate gradient (CG). This iterative solver requires only matrix-vector products, never the full matrix.
-
Estimate the diagonal via the Rademacher trick. The Rademacher diagonal estimator averages the outputs of random “probe” vectors fired through the system, producing an unbiased estimate of the diagonal of the inverse covariance. An O(D²) impossibility becomes a statistical estimate that converges reliably.
-
Parallelize across GPUs. The K linear systems are independent, so they run simultaneously on GPU hardware. Most SBL competitors can’t exploit this because of memory constraints.
Per-iteration cost drops from O(D³) to O(UKτD), where τD is one matrix-vector multiply and U, K are small fixed hyperparameters. Memory drops from O(D²) to O(DK).
For measurement matrices satisfying the restricted isometry property (RIP), a standard condition in compressed sensing that ensures measurements capture the signal broadly, the authors prove U and K can stay fixed as D grows. More unknowns doesn’t mean more probes.

In simulations, CoFEM matches standard EM on sparse signal recovery while cutting runtime by hundreds to thousands of times. With GPU acceleration, computation that previously took hours shrinks to seconds.
Why It Matters
SBL’s ability to quantify uncertainty has always set it apart from LASSO and ℓ₁-regularized compressed sensing. But that ability was locked behind a computational wall for any problem at scale. CoFEM removes that wall.
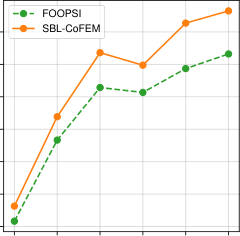
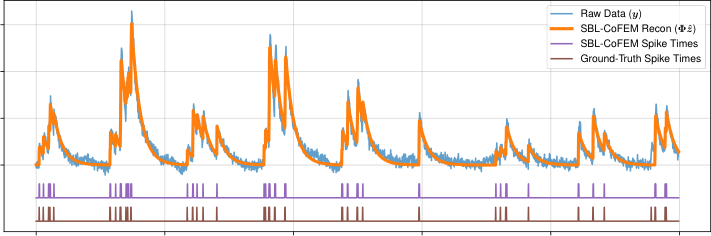
The paper tests this on two real applications. In calcium imaging deconvolution, the goal is to infer when neurons fire from fluorescence traces. CoFEM-equipped SBL recovers firing patterns with competitive accuracy while delivering uncertainty estimates that purely deterministic methods cannot provide.
In multi-contrast MRI reconstruction, the task is reconstructing multiple tissue-contrast images simultaneously from undersampled k-space data. CoFEM handles volumetric brain scans in practical compute times.
Both extensions (multi-task learning across MRI contrasts, non-negativity constraints for calcium signals) slot into the CoFEM framework without difficulty.
There’s a broader lesson here. Many Bayesian methods remain confined to low-dimensional settings because their inference costs scale with the cube of the problem size. CoFEM charts a way out: figure out what you actually need from an expensive intermediate object, then estimate it cheaply with randomized numerical linear algebra. The same strategy applies anywhere uncertainty quantification matters, but brute-force Bayesian computation won’t fit in memory or time budgets.
Bottom Line: CoFEM makes Sparse Bayesian Learning fast enough for high-dimensional real-world problems by replacing explicit covariance matrix inversion with randomized linear solvers. The result is thousandfold speedups with no accuracy loss, opening the door to principled uncertainty quantification at scale.
IAIFI Research Highlights

This work sits at the intersection of Bayesian statistics, numerical linear algebra, and signal processing. The method applies to scientific imaging problems (MRI reconstruction, neural calcium imaging) where uncertainty estimates carry real interpretive weight.

CoFEM shows that randomized numerical methods can accelerate Bayesian inference from cubic to near-linear complexity in the covariance-matrix EM setting, substantially widening what probabilistic machine learning can tackle in practice.

By making principled uncertainty quantification tractable in high-dimensional inverse problems, CoFEM provides a practical tool for physics-adjacent domains whose mathematical structure mirrors sensing problems in fundamental physics.
Natural next steps include extending CoFEM to non-Gaussian priors and to large-scale inverse problems in astrophysics and particle physics; see [arXiv:2105.10439](https://arxiv.org/abs/2105.10439) for the full technical treatment.

